Virus-host cell interactions
The adhesion and internalization of viral particles by cells is a mechanical process. We have used AFM-based, single-virus force spectroscopy to characterize the interaction between viruses and host cells. This approach enables the extraction of the binding properties, such as the on- and off- rates, binding valency, and affinity alterations, of viral surface-host receptor interactions at the single-virion level. By combining with fluorescence microscopy, the technique offers a powerful tool to determine the dynamic mechanical process of viral-host membrane fusion and viral internalization and to allow the screening of viral entry blockers. This research theme concerns the cellular invasion of two emerging viruses: Ebola and SARS-CoV-2.
The most recent Ebola virus (EBOV) outbreak posed a significant threat to the world. My ongoing project concerns how the filamentous EBOV interacts on the host cell surface with its receptor, the T-cell immunoglobulin and mucin domain (TIM) family proteins. My group used single-molecule atomic force microscopy and biomechanical models to quantify the specific interaction forces between TIM and EBOV pseudovirions (nonpathogenic model systems). These findings may help develop a novel antiviral therapy to prevent EBOV entry without compromising TIM’s physiological role in taking up apoptotic cell debris. The EBOV research is currently supported by an NIH R15 grant and an NSF grant.
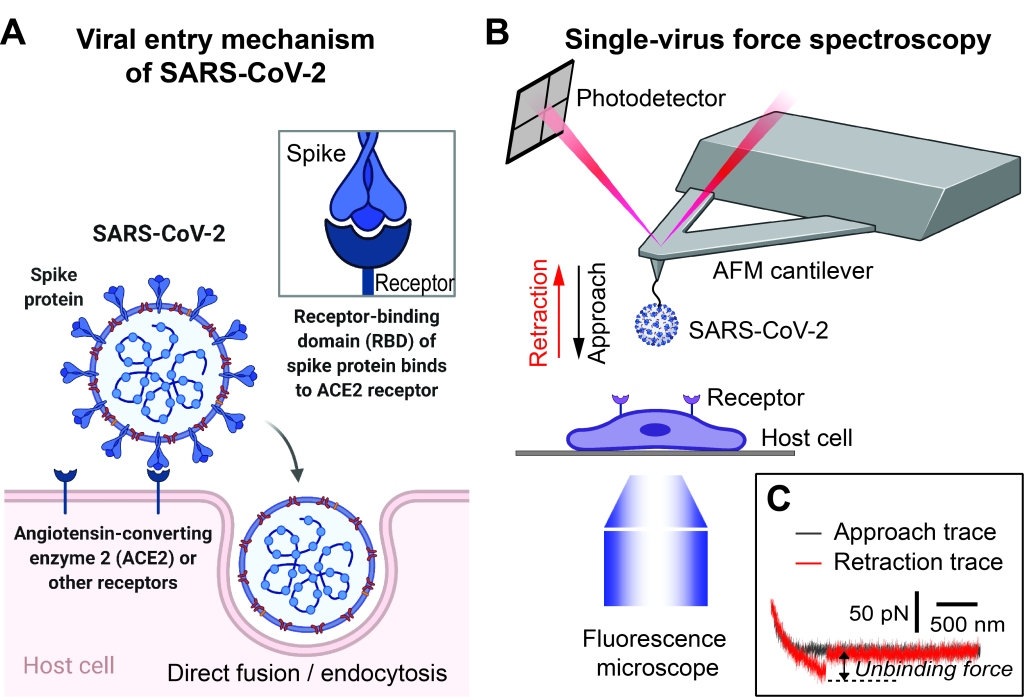
The current coronavirus disease-2019 (COVID-19) pandemic has had devastating impacts across the world. SARS-CoV-2, the virus causing COVID-19, uses its surface spike (S) protein to attach to host cell surface receptors to enter human host cells (Fig. A). Besides the best-known receptor, ACE2, several cell-surface molecules have been reported to bind to S protein and help mediate SARS-CoV-2 entry. Our preliminary studies using single-virus force spectroscopy (Fig. B,C) consistently show that these molecules can bind to the SARS-CoV-2 S protein with comparable affinities to those of ACE2. Therefore, the possible multiple receptor utilization could partially explain the broad tissue tropism and systemic complications of the SARS-CoV-2 infection. However, it remains puzzling how the S protein can bind to these structurally diverse molecules with high affinity. Additionally, recent structural data shows that most of the S protein surface is covered by glycans; it can only bind to a receptor without glycan interference when the S protein’s receptor-binding domain (RBD) is in the “up position.” Therefore, we hypothesize that limited regions on the S protein that are not covered by glycans, including the RBD in the up position and the S1/S2 junctional region, may be responsible for binding all the receptors. We are currently testing the hypothesis using combined approaches of single-virus force spectroscopy, single-virus fluorescence imaging, all-atom molecular modeling and simulation, and pseudovirus internalization/entry assays. Moreover, SARS-CoV-2 has two entry routes (direct viral-host membrane fusion or endocytic/macropinocytic internalization followed by endosomal entry) (Fig. A), and the exact entry pathways mediated by these receptors are not yet clear. We are currently determining whether every individual S protein-receptor interaction is more prone to mediating direct viral-host membrane fusion or viral endocytic/macropinocytic internalization. The study will elucidate the structural and biophysical mechanisms behind the S protein’s receptor recognition and utilization and allow us to identify new targets for antiviral therapies to treat COVID-19’s systemic complications.
In addition, we are collaborating with a group of Lehigh faculties (Xuanhong Cheng, Himanshu Jain, Anand Jagota, and Yaling Liu) and external collaborators (Kavssery Ananthapadmanabhan of University of Cincinnati, and Solvay USA Inc.) to develop long-lasting virucidal coatings against enveloped viruses including SARS-CoV-2. The work is supported by the Commonwealth of Pennsylvania.
Lymphocyte adhesion mediated by integrins
Integrin is a heterodimeric membrane receptor that is expressed on most eukaryotic cells and serves as an adhesion receptor and mechanical sensor to transmit signals bidirectionally across the plasma membrane. Using atomic force microscopy (AFM), we examined an allosteric regulatory model through which the ligand-binding affinity can be increased rapidly when a force is allowed to stretch the C-terminal of integrin’s ligand-binding I-domain. This type of mechanism may account for the rapid ligand affinity adjustment during leukocyte migration. More recently, we studied how integrin-ligand bond lifetime and mechanical properties are regulated on lymphocytes by chemokine signals.
Representative publications:
- Cao, W., Kim, S., Dong, C., Hou, D., Im, W. and Zhang, X.F.* (2021): Biomechanical Characterization of SARS-CoV-2 Spike RBD and Human ACE2 Protein-Protein Interaction. Biophysical Journal, 120:1211-1219.
- Cui, X., Lapinski, N., Zhang, X.F., and Jagota, A. (2021): Length of Mucin-Like Domains Enhance Cell-Ebola Virus Adhesion by Increasing Binding Probability. Biophysical Journal, 120:781-790.
- Wang, J., Lapinski, N., Zhang, X.F., and Jagota, A. (2020): Adhesive Contact Between Cylindrical (Ebola) and Spherical (SARS-CoV-2) Viral Particles and a Cell Membrane. Mechanics of Soft Materials, 2 (1), 1-9.
- Dragovich, M.A., Fortoul, N., Jagota, A., Schutt, K., Xu, Y., Sanabria, M., Moller-Tank, S., Maury, W., and X.F.* (2019): Biomechanical characterization of TIM protein-mediated Ebola virus-host cell adhesion. Scientific Reports, 9:267.
- Bondu, V., Wu, C., Cao, W., Simons, P., Gillette, J., Zhu, J., Erb, L., Zhang, X.F.* and Buranda, T.* (2017): Low affinity binding in cis to P2Y2R mediates force-dependent integrin activation during hantavirus infection. Molecular Biology of the Cell, 28(21):2887-2903.
- Wang, S., Wu, C., Zhang, Y., Zhong, Q., Sun, H., Cao, W., Ge, G.X., Li, G.H., Zhang, X.F.*, and Chen, J.F.* (2018): The Structural basis of chemokine-mediated switch in integrin α4β7 ligand specificity. The Journal of Cell Biology, 217 (8): 2799-2812.
- Fu, X. Xu, Y., Wu, C., Moy, V.T., and Zhang, X.F.* (2015): Anchorage-dependent binding of integrin I-domain to adhesion ligands. Journal of Molecular Recognition, 28:385-92.
- Cao, Y., Hoeppner, L.H., Bach, S., Guangqi, E., Guo, Y., Wang, E., Wu, J., Cowley, M.J., Chang, D.K., Waddell, N., Grimmond, S.M., Biankin, A.V., Daly, R.J., Zhang, X.F., and Mukhopadhyay, D. (2013): Neuropilin-2 promotes extravasation and metastasis by interacting with endothelial a5 integrin. Cancer Research, 73(14):4579-4590.
- Rico, F., Zhang, X., and Moy, V.T. (2011): Probing cellular adhesion at the single molecular level. Chapter 11 (p. 225-262) in Life at the Nanoscale: Atomic Force Microscopy of Live Cells (Yves Dufrêne, Ed.), Pan Stanford Publishing Ltd.
- Zhang, X.*, Rico F., Xu, A.J., and Moy, V.T. (2009): Atomic force microscopy of protein-protein interactions. Chapter 19 in Handbook of Single-Molecule Biophysics (Peter Hinterdorfer and Antoine van Oijen, Ed. ISBN: 978-0-387-76496-2), Springer Publishing(http://www.springerlink.com/content/q85x87x114735130/).
- Zhang, X. and Moy, V.T. (2006): Intermolecular forces of leukocyte adhesion molecules. Chapter 9 (p. 159-168) in Force Microscopy: Application in Biology and Medicine (Bhanu P. Jena and Johann Karl H. Horber, Ed.), John Wiley & Sons Inc. Press.
- Zhang, X.*, Wojcikiewicz, E., and Moy, V.T (2006): Dynamic adhesion of T-lymphocytes to endothelial cells revealed by atomic force microscopy. Experimental Biology and Medicine, 231: 1306-12.
- Zhang, X., Craig, S.E., Kirby, H., Humphries, M.J. and Moy, V.T. (2004): Molecule basis for the dynamic strength of the integrin a4β1/VCAM-1 interaction. Biophysical Journal, 87: 3470-3478.
- Zhang, X., Chen, A., DeLeon, D., Li, H., Noiri, E., Moy, V.T. and Goligorsky, M.S. (2004): Atomic force microscopy measurement of leukocyte-endothelial interaction. American Journal of Physiology-Heart and Circulatory Physiology, 286: H359-367.
Single-virus force spectroscopy. (A) Schematic of a SARS-CoV-2 viral particle and its entry mechanism. (B) Schematic of our AFM-based, single-virus force spectroscopy experimental platform. (C) Representative AFM force-distance traces of a SARS-CoV-2 pseudovirus-functionalized tip interacting with a live host cell.